Morphological characteristics and antibacterial activities of Phi1 and Phi3
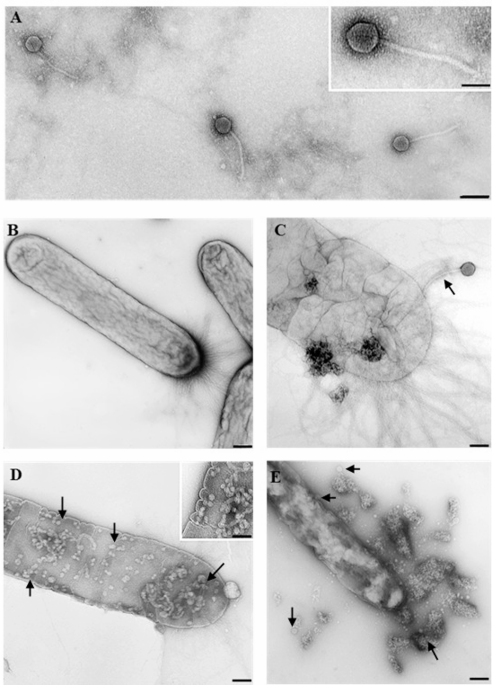
TEM analysis showed that phage Phi3 exhibits morphological characteristics typical of the siphovirus morphotype, with an isometric head approximately 60 nm in diameter, and a long, flexible, non-contractile tail of 225 ± 10 nm in length (Fig. 1A). In contrast, phage Phi1 has the morphology of a podovirus, with an isometric head measuring 60 ± 3 nm and a short stubby tail approximately 10–15 nm in length (Fig. 2A). Furthermore, interactions between phage Phi1 and Phi3 and Xfp cells were examined. Micrographs revealed multiple stages of the infection cycle for both phages, including phage adsorption to the bacterial cell surface (Figs. 1C and 2B), phage replication inside the Xfp cells (Figs. 1D and 2C) and progressive disruption of the Xfp cell membrane (Figs. 1E and 2D), indicating efficient phage infection. In plaque assays using Xcc as the host, phage Phi1 produced large, clear plaques, whereas phage Phi3 generated small, clear plaques, with no halo formation observed for either phage (Fig. 3).
Fig. 1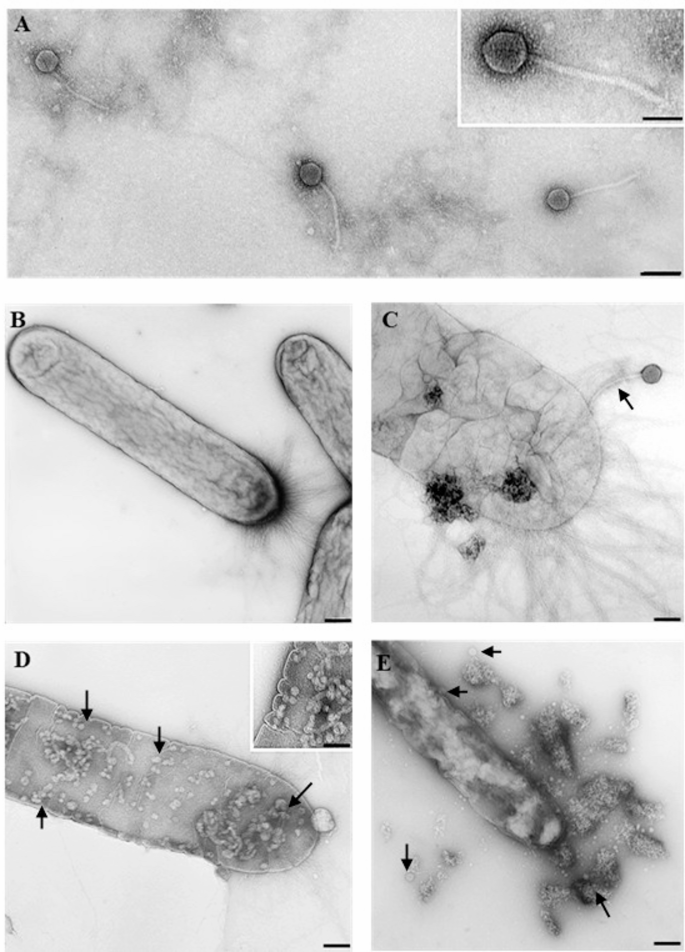
Transmission electron micrographs showing morphology of phage Phi3 and the infection cycle on Xylella fastidiosa subsp. pauca cells. (A) Phi3 particles resembling the siphovirus morphotype (B) Untreated Xfp cells, used as control. (C, D) Phi3-treated Xfp cells showing adsorption (arrow) and replication of Phi3 (arrowhead). (E) Lysis of Phi3-treated Xfp cell. Bar = A, B, C, D, E 100 nm, insets 50 nm.
Fig. 2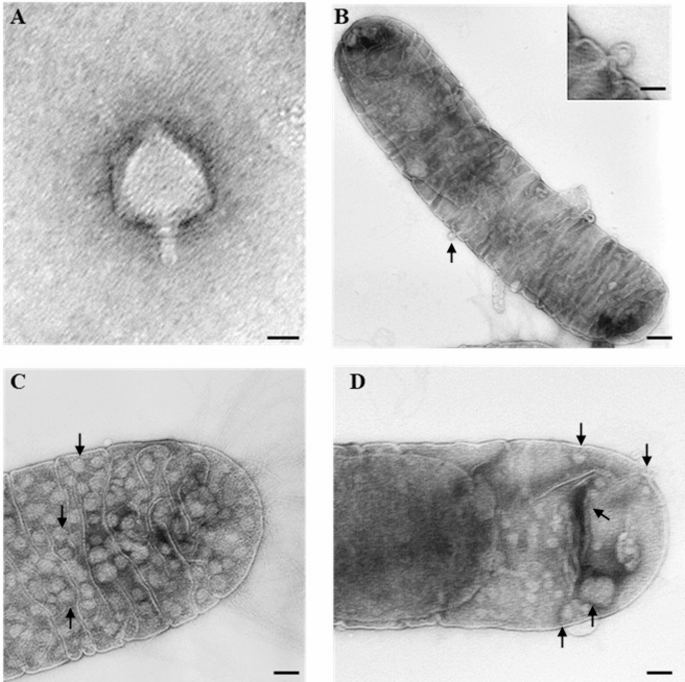
Transmission electron micrographs showing morphology of phage Ph1 and the infection cycles on Xylella fastidiosa subsp. pauca cells. (A) Phi1 particle resembling the podovirus morphotype. (B, C) Phi1-treated Xfp cells showing adsorption (arrow) and replication of Phi1 (arrows). (D) Phi1-treated Xfp cell showing membrane alteration. Bar = A 25 nm; B, C 50 nm, inset 25 nm; D 150 nm).
Fig. 3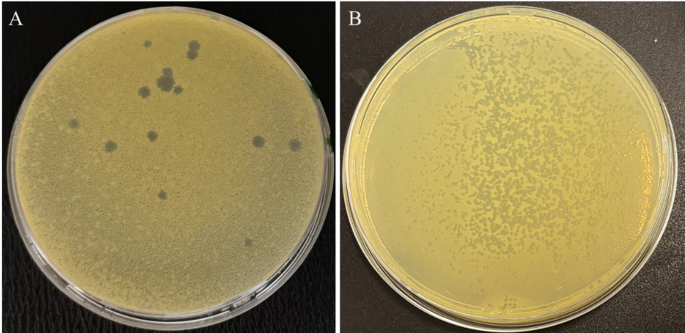
Plaque morphology of bacteriophages Phi1 (A) and Phi3 (B) on double-layer agar plates using Xanthomonas campestris pv. campestris as the host strain.
Genomic and phylogenetic analyses of Phi1 and Phi3
The complete genomes of Phi1 and Phi3 were sequenced using the Illumina HiSeq 2500 platform. After quality trimming, adapter removal, read merging, error correction, normalization, and duplicate removal, 2,519 and 3,351 high-quality reads were retained for Phi1 and Phi3, respectively, corresponding to approximately 18.8 Mb and 15.1 Mb of raw sequence data. De novo assembly generated single contigs for each genome, with finalized genome sizes of 44,345 bp for Phi1 and 55,413 bp for Phi3. The estimated sequencing coverage was approximately 425× for Phi1 and 271× for Phi3, calculated as the total number of sequenced bases divided by the genome length. The N50 values are equivalent to the genome sizes, confirming that the assemblies are complete and high quality. The complete genome of Phi1 consisted of 62 coding sequences (CDS), of which the functions of 26 CDS (41.9%) were predicted; whereas the complete genome of Phi3 comprised 75 CDS, of which functional annotations were assigned to 34 (45.3%). No tRNA genes were identified in either phage genome based on annotation using the Pharokka pipeline. The genomic characteristics and genome maps of phages Phi1 and Phi3 are shown in Table 2; Fig. 4. PhaTYP analyses predicted that both phages undergo a strictly lytic infection cycle, with no evidence of temperate markers, virulence factors or AMR genes. These findings were corroborated by ABRICATE and AMRFinderPlus results, which confirmed the absence of AMR and virulence genes in Phi1 and Phi3 genomes. These results indicate that Phi1 and Phi3 could be suitable for use as biocontrol agents.
Comparative genomic analysis using BLASTn against the NCBI nucleotide database revealed that Phi1 and Phi3 are closely related to phages in the Pradovirus genus (Autonoeviridae family) and the Salvovirus genus (Casjensviridae family), respectively. Both genera, Pradovirus and Salvovirus, comprise phages known to infect multiple species within the genera Xylella and Xanthomonas, including Xcc and Xfp. Phi1 shares the highest sequence similarity (identity: 92.67%, coverage: 93%) with Xanthomonas phage phi Xc10 (GenBank: NC_047840), while Phi3 showed the closest homology to Xylella phage Salvo (GenBank: NC_042345), with 98.44% identity and 91% coverage. The comparative genomic alignments between Phi1 and Phi3 and their respective reference genomes are illustrated in Fig. 5. Additionally, ANI analysis using the OrthoANIu algorithm yielded a value of 92.52% between Phi1 and Xanthomonas phage phi Xc10, suggesting that Phi1 represents a novel phage species, as it falls below the established 95% species demarcation threshold of the “International Committee on Taxonomy of Viruses” (ICTV). In contrast, phage Phi3 exhibited an OrthoANIu value of 96.70% when compared to Xylella phage Salvo, indicating that it is likely the same species. Furthermore, phylogenetic analysis based on whole-proteome comparisons, generated using ViPTree, corroborated the genomic analyses and positioned Phi1 and Phi3 alongside other Xylella and Xanthomonas phages within the genera Pradovirus and Salvovirus, respectively (Fig. 6). The complete genome sequences of Phi1 and Phi3 were deposited in in NCBI GenBank with the accession numbers PX095379 and PX095380, respectively.
Table 2 Comparative genomic features of phages Phi1, Phi3, and their closest GenBank-reported relatives.Fig. 4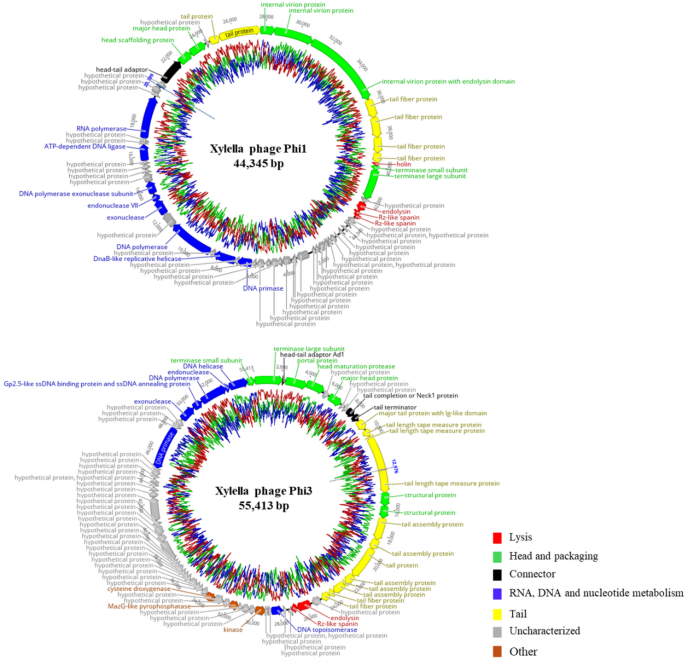
Circular genome maps of Phi1 and Phi3 generated using Geneious Prime 2025.1.1. The inner rings illustrate the GC content percentage, while predicted coding sequences identified by Pharokka are depicted as coloured arrows, with coloration reflecting their assigned functional categories.
Fig. 5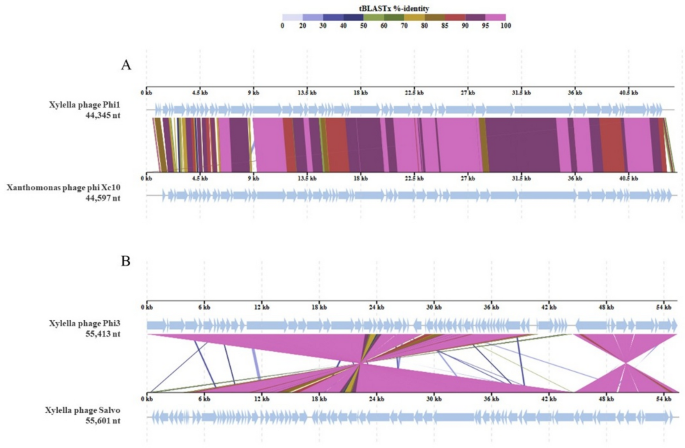
Comparative genomic alignments between Phi1 (A) and Phi3 (B) and their respective most similar genomes, generated using ViPTree. The colored vertical blocks between the genomes indicate levels of nucleotide similarity.
Fig. 6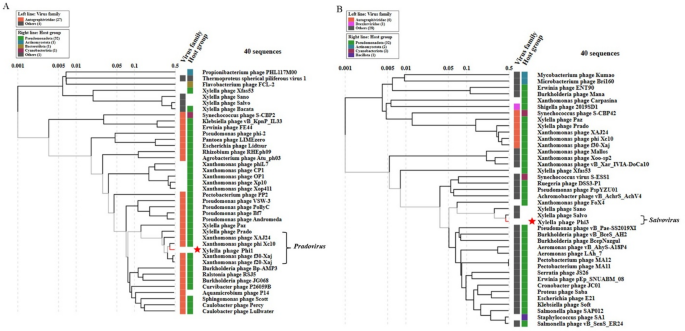
Proteomic trees of Phi1 (A) and Phi3 (B), generated by ViPTree based on genome-wide sequence similarities computed by tBLASTx, showing the allocation of Phi1 and Phi3 among species belonging to Pradovirus and Salvovirus, respectively.
In silico identification of putative antibiofilm-associated depolymerases in Phi1 and Phi3
Computational identification of depolymerase-encoding genes within the genomes of Phi1 and Phi3 was conducted using DePP. This analysis resulted in the prediction of three proteins (P19, P44, and P46) with high probabilities (92–98%) of depolymerase activity, whose predicted structural and physicochemical characteristics are summarized in Table 3. The three proteins range from 787 to 834 amino acids (85–91 kDa) and exhibit mildly acidic theoretical pI values (4.6–5.3). ProtParam analysis classified the three putative depolymerases as hydrophilic (negative GRAVY values) and stable (instability index < 40), with half-lives > 10 h in E. coli, supporting their suitability for recombinant expression. Subsequent HHpred analysis of the three proteins initially predicted as depolymerases by DePP indicated that P19 and P44 exhibit homology to phage tail-associated structural proteins, without detectable similarity to any characterized depolymerases, while P46 was assigned (100% probability) to the peptidoglycan hydrolase gp36 (Table 3). Peptidoglycan hydrolase gp36 is an exolysin that catalyzes the cleavage of the host peptidoglycans, playing a role in ejection of the phage DNA into the host cell at the initiation of infection35. Additionally, the (trans)glycosylase activity of recombinant gp36C has been experimentally validated against Escherichia coli WK6 and Pseudomonas aeruginosa AO135. This strongly suggests that P46 functions as a peptidoglycan hydrolase rather than exopolysaccharide depolymerase. However, experimental biochemical validation is required to confirm the specific enzymatic activities of the three proteins.
Table 3 Predicted phage-encoded depolymerases and their physicochemical and structural properties derived from DePP, ProtParam, and HHpred analyses.Predicted lysis modules of Phi1 and Phi3
Genomic annotation using Pharokka, with subsequent structural validation through HHpred analysis, enabled prediction of the lysis module organization in Phi1 and Phi3 (Fig. 3; Table 4). In Phi1, the lysis module consists of four genes encoding for a 66 amino acid holin (CDS_0052) with predicted membrane-pore forming function supported by 84.45% homology to HOLIN_BPT7; a 184 amino acid endolysin (CDS_0056) exhibiting > 99% probability matches to the peptidoglycan-degrading ENLYS_BPB03; and a pair of Rz-like spanins (CDS_0057/CDS_0058) (Table 4). Phi3 encodes a two-gene lysis module consisting of a 345 amino acid endolysin (CDS_0026, 98.74% match to ENLYS_BPB03) and a 147 amino acid spanin (CDS_0027, 95.62% match to SPAN1_BPMU) (Table 4). The high-confidence HHpred matches (> 84% probability) strongly supports the functional annotations of the predicted lysis-associated proteins, which may represent promising candidates for synergistic applications in the management of Xf infections in the future.
Table 4 Predicted lysis-associated proteins of Phi1 and Phi3 and their physicochemical properties.Off-target effects of Phi1 and Phi3
A total of fourteen bacterial isolates belonging to the genera Paenibacillus, Peribacillus, Bacillus, and Neobacillus were obtained from tissues of olive trees (Table 5). These isolates, together with the beneficial bacterial strains listed in Table 1, were used to assess the off-target effects and suitability of Phi1 and Phi3 as biocontrol agents. Results showed that neither Phi1 nor Phi3 demonstrated lytic activity against any of the tested non-target bacteria, thereby indicating a restricted host range and suggesting a minimal threat to the indigenous olive-associated microbiota. This pronounced degree of specificity reinforces their prospective utilization as safe and targeted biocontrol agents against Xfp in olive agricultural practices. Furthermore, the taxonomic diversity of the isolated bacteria offers valuable insights into the bacteriome associated with olive trees in the Apulia region, serving as a foundational reference for subsequent investigations concerning olive-associated microbial communities and identification of potential biocontrol agents against Xf.
Table 5 Bacterial isolates recovered from olive trees for off-target effects assessment of Phi1 and Phi3.Temperature and pH stability
The stability of Phi1 and Phi3 under a range of thermal and pH conditions was evaluated. Thermal tolerance assays indicated that both phages maintained infectivity across a large temperature spectrum, spanning from − 20 °C to 60 °C. Nevertheless, a total loss of biological viability was observed at a temperature of 70 °C, indicating a significant thermal threshold (Fig. 7). Simultaneously, the stability of pH was assessed by subjecting phage suspensions to incubation in phage buffer with pH levels ranging from 3 to 10. The results revealed that infectivity was maintained within the pH spectrum of 3 to 10 (Fig. 7). These findings imply that the phages exhibited significant tolerance to extreme physicochemical environments, thereby reinforcing their prospective application in diverse biotechnological and agricultural contexts. All statistical analyses were performed on log₁₀-transformed PFU/mL values, as titer data follow a log-normal distribution. On this scale, some conditions that appear far apart on the linear axis (e.g., 10³ vs. 10⁸ PFU/mL or 10⁶ vs. 10⁹ PFU/mL) show overlapping variance (Fig. 7), and with a small sample size (n = 3) these differences do not reach significance in the Dunn’s test. Thus, the grouping letters indicate non-significant differences on the log scale, not equivalence of absolute PFU values, but the “not statistically different” in this context does not imply biological similarity, but rather that the within-group variability relative to sample size limits statistical discrimination despite the visually large differences in raw titers.
Fig. 7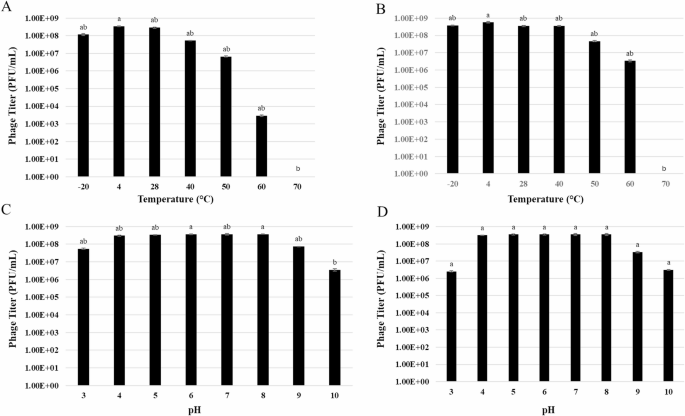
Histograms showing pH and thermal stability tests of Phi1 and Phi3. (A, C) Phi1 stability after exposure to different temperatures and pH values for 24 h. (B, D) Phi3 stability after exposure to different temperatures and pH values for 24 h. Phage titers were determined using double agar overlay method. Data represent the mean ± standard deviation of three biological replicates (n = 3). Different letters (a-b) above the bars indicate statistically significant differences (p < 0.05) between treatments.

Dining and Cooking